[1]:
%matplotlib widget
3) Complete spectrum analysis
In this example, we shall fit all the emission lines on the spectrum of the Green Pea galaxy GP121903 (see Fernandez et al 2021) using the Spectrum.fit.frame. This function can fit multiple lines from an input lines frame. Moreover, we are going to show the recommended 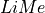 workflow: Using external files with the line bands and fitting configuration:
workflow: Using external files with the line bands and fitting configuration:
In this exercise, we are going to follow the recommended
You can download this spectrum from the examples/sample_data. This tutorial can found as a script and a notebook on the examples folder.
Loading the data
Let’s start by importing the script packages and declaring the data location:
[2]:
import numpy as np
from astropy.io import fits
from IPython.display import Image, display
from pathlib import Path
import lime
[3]:
# State the input files
obsFitsFile = '../sample_data/spectra/gp121903_osiris.fits'
lineBandsFile = '../sample_data/osiris_bands.txt'
cfgFile = '../sample_data/osiris.toml'
# Spectrum parameters
z_obj = 0.19531
norm_flux = 1e-18
The configuration file (cfgFile) can be read with the .load_cfg function:
[4]:
# Load configuration
obs_cfg = lime.load_cfg(cfgFile)
# Get object redshfit and normalization flux
z_obj = obs_cfg['sample_data']['z_array'][2]
norm_flux = obs_cfg['sample_data']['norm_flux']
This is a .toml configuration text file which is loaded as a dictionary of dictionaries.
[5]:
import pprint
pprint.pprint(obs_cfg)
{'default_line_fitting': {'Ar4_4711A_m': 'Ar4_4711A+He1_4713A',
'H1_3889A_m': 'H1_3889A+He1_3889A',
'O2_3726A_m': 'O2_3726A+O2_3729A',
'O2_7319A_m': 'O2_7319A+O2_7330A'},
'gp121903_line_fitting': {'H1_6563A_b': 'H1_6563A+N2_6584A+N2_6548A',
'N2_6548A_amp': {'expr': 'N2_6584A_amp/2.94'},
'N2_6548A_kinem': 'N2_6584A',
'O1_6300A_b': 'O1_6300A+S3_6312A',
'O3_5007A_b': 'O3_5007A+O3_5007A_k-1',
'O3_5007A_k-1_amp': {'expr': '<100.0*O3_5007A_amp',
'min': 0.0},
'O3_5007A_k-1_sigma': {'expr': '>2.0*O3_5007A_sigma'},
'S2_6716A_b': 'S2_6716A+S2_6731A',
'S2_6731A_kinem': 'S2_6716A'},
'sample_data': {'files_list': ['gp030321_BR.fits',
'gp101157_BR.fits',
'gp121903_BR.fits'],
'norm_flux': 1e-17,
'object_list': ['gp030321', 'gp101157', 'gp121903'],
'zErr_array': [7.389e-05, 0.000129, 0.0001403],
'z_array': [0.16465, 0.14334, 0.19531]}}
You can read more about how to use these files on the fitting configuration documentation
You can load the bands file from the second tutorial as a pandas dataframe using the load_frame function:
[6]:
# Load line bands
bands = lime.load_frame(lineBandsFile)
In the first tutorial, we opened the observation using the standard aproach with astropy. This time we are going to use 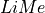 functions to read a .fits file:
functions to read a .fits file:
[7]:
gp_spec = lime.Spectrum.from_file(obsFitsFile, instrument='osiris', redshift=z_obj, norm_flux=norm_flux)
gp_spec.plot.spectrum(rest_frame=True)
Line detection
In the previous tutorial, we manually selected the bands, so we are certain that all the lines are present. However in large datasets it may not be possible to inspect all the lines manually. You can use 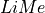 functions to help you with this task.
functions to help you with this task.
The current method available is based on an intensity threshold for the emission or absorption lines. To improve the performance fo this task, the first step consists in adjusting the observation continuum:
[8]:
gp_spec.fit.continuum(degree_list=[3, 6, 6], emis_threshold=[3, 2, 1.5], plot_steps=True)
The task Spectrum.fit.continuum provides an interactive analysis: The user introduces a list of degrees for the polynomial degrees and a multiplicative factor in the degree_list and emis_threshold, respectively. At each step, the algorihm fits a polinomial of the given order, rejecting the points which are above/below the multiplicative factor of the standard deviation of the residual betwen the observed spectrum and the fit continuum. The fitting excludes all the points which were
rejected from the previous iterations. If the user does not include an abs_threshold array, the algorithm uses the multiplicative factor from the emis_threshold.
Once we have computed the continuum we can use the Spectrum.line_detection function to detect peaks or trough and compare them against the theoretical line locations:
[9]:
# Find the lines
matched_bands = gp_spec.line_detection(lineBandsFile, sigma_threshold=3, plot_steps=True)
In this function, the sigma_threshold argument states the factor above the continuum sigma flux threshold for a line detection. The argument emis_type=True for emission lines and emis_type=False for absorption lines.
The user needs to include a bands dataframe, such as the lineBandsFile we computed from the previous tutorial.
You can plot a set of bands in you spectrum with the line_bands argument in the .plot.spectrum function:
[10]:
gp_spec.plot.spectrum(label='GP121903 matched lines', line_bands=matched_bands, log_scale=True)
Line measurement
Now we are going to measure all the lines in this bands dataframe using the Spectrum.fit.frame function. This function will read the obj_bands_file and proceed to fit all the lines taking into consideration the lines fitting configuration from the obs_cfg dictionary.
[11]:
gp_spec.fit.frame(bands=matched_bands, fit_conf=obs_cfg, id_conf_prefix='gp121903')
Line fitting progress:
[==========] 100% of 27 lines (O2_7319A_m)
Please remember: You can introduce the line bands and fitting configuration fit_conf file addresses and the .fit. functions will read them for you.
The Spectrum.fit.frame function allows the user to combine a “default” fitting configuration with an “individual” fitting configuration. The former configuration parameters are specified with default_conf_prefix argument, while the latter uses the id_conf_prefix. For example, in the previous fitting, we had the “default_line_fitting” and the “gp121903_line_fitting” form the configuration file:
[12]:
print(f'# Default line fitting')
pprint.pprint(obs_cfg['default_line_fitting'])
print(f'\n# Individual line fitting')
pprint.pprint(obs_cfg['gp121903_line_fitting'])
# Default line fitting
{'Ar4_4711A_m': 'Ar4_4711A+He1_4713A',
'H1_3889A_m': 'H1_3889A+He1_3889A',
'O2_3726A_m': 'O2_3726A+O2_3729A',
'O2_7319A_m': 'O2_7319A+O2_7330A'}
# Individual line fitting
{'H1_6563A_b': 'H1_6563A+N2_6584A+N2_6548A',
'N2_6548A_amp': {'expr': 'N2_6584A_amp/2.94'},
'N2_6548A_kinem': 'N2_6584A',
'O1_6300A_b': 'O1_6300A+S3_6312A',
'O3_5007A_b': 'O3_5007A+O3_5007A_k-1',
'O3_5007A_k-1_amp': {'expr': '<100.0*O3_5007A_amp', 'min': 0.0},
'O3_5007A_k-1_sigma': {'expr': '>2.0*O3_5007A_sigma'},
'S2_6716A_b': 'S2_6716A+S2_6731A',
'S2_6731A_kinem': 'S2_6716A'}
Please remember: The Spectrum.fit.frame function updates the default parameters with the new ones. This means that only the common entries in the default configurations are replaced. Consequently, in the fittings of GP121903 the configuration used would be:
[13]:
pprint.pprint({**obs_cfg['default_line_fitting'], **obs_cfg['gp121903_line_fitting']})
{'Ar4_4711A_m': 'Ar4_4711A+He1_4713A',
'H1_3889A_m': 'H1_3889A+He1_3889A',
'H1_6563A_b': 'H1_6563A+N2_6584A+N2_6548A',
'N2_6548A_amp': {'expr': 'N2_6584A_amp/2.94'},
'N2_6548A_kinem': 'N2_6584A',
'O1_6300A_b': 'O1_6300A+S3_6312A',
'O2_3726A_m': 'O2_3726A+O2_3729A',
'O2_7319A_m': 'O2_7319A+O2_7330A',
'O3_5007A_b': 'O3_5007A+O3_5007A_k-1',
'O3_5007A_k-1_amp': {'expr': '<100.0*O3_5007A_amp', 'min': 0.0},
'O3_5007A_k-1_sigma': {'expr': '>2.0*O3_5007A_sigma'},
'S2_6716A_b': 'S2_6716A+S2_6731A',
'S2_6731A_kinem': 'S2_6716A'}
Plotting and saving the measurements
The fitted profiles can be over-plotted on the input spectrum setting the include_fits=True parameter on the Spectrum.plot.spectrum function
[14]:
# Display the fits on the spectrum
gp_spec.plot.spectrum(include_fits=True, rest_frame=True)
Additionally, you can also plot the results as a grid using the Spectrum.plot.grid
[15]:
# Display a grid with the fits
gp_spec.plot.grid()
You can plot individual line fittings with the Spectrum.plot.band function:
[16]:
gp_spec.plot.bands('H1_6563A')
Finally, we save the tabulated measurements with:
[17]:
# Save the data
gp_spec.save_frame('../sample_data/example3_linelog.fits', page='GP121903_a')
or
[18]:
lime.save_frame('../sample_data/example3_linelog.fits', gp_spec.frame, page='GP121903b')